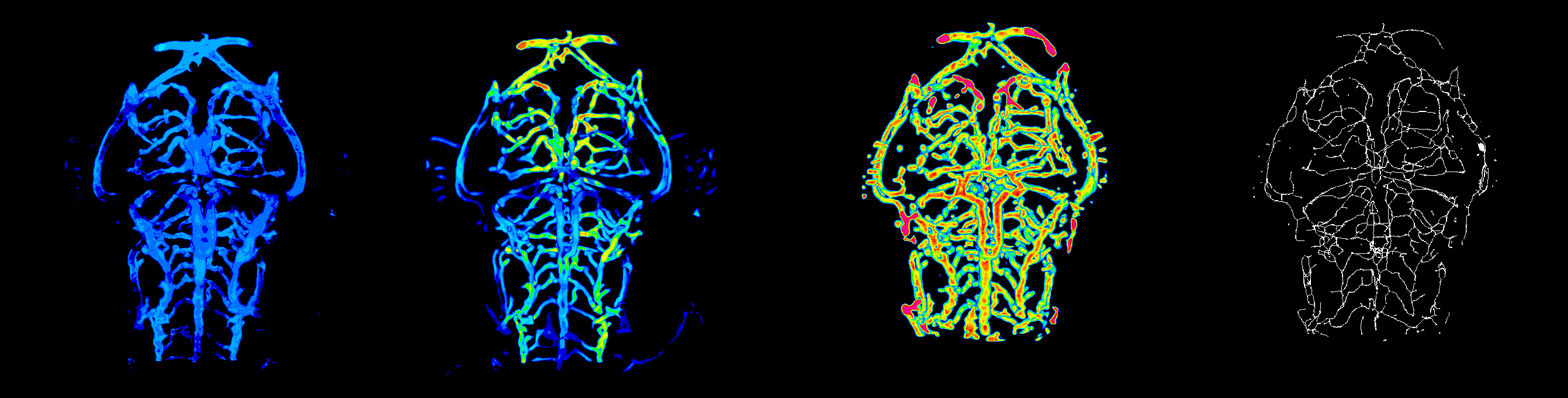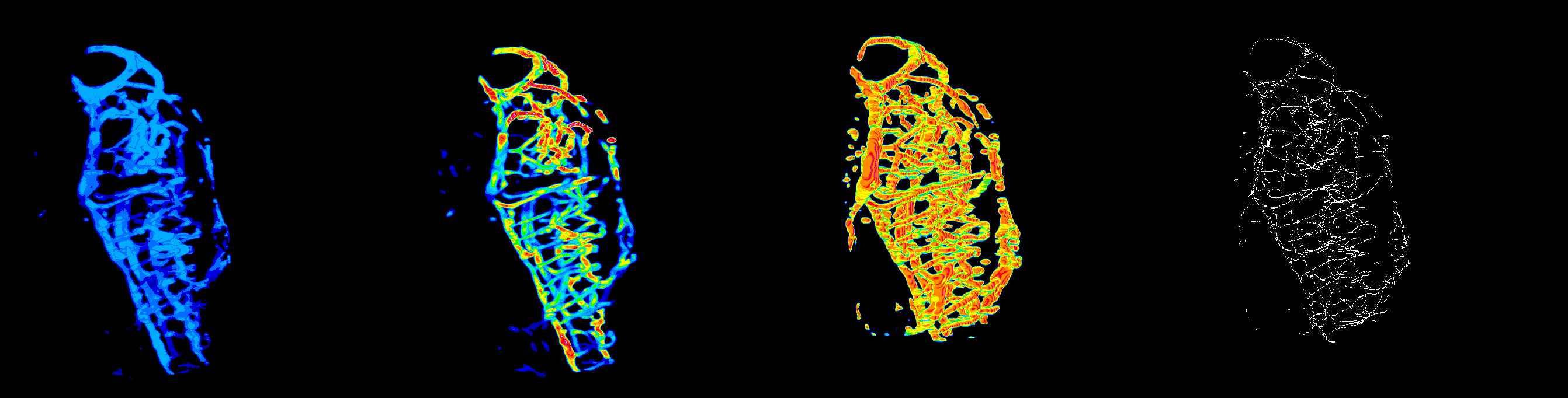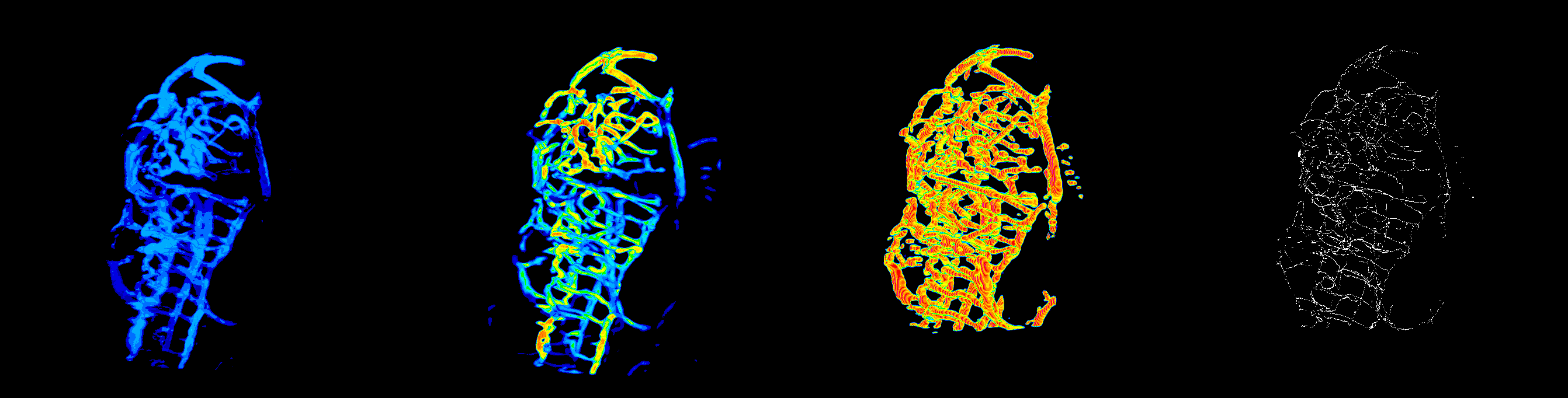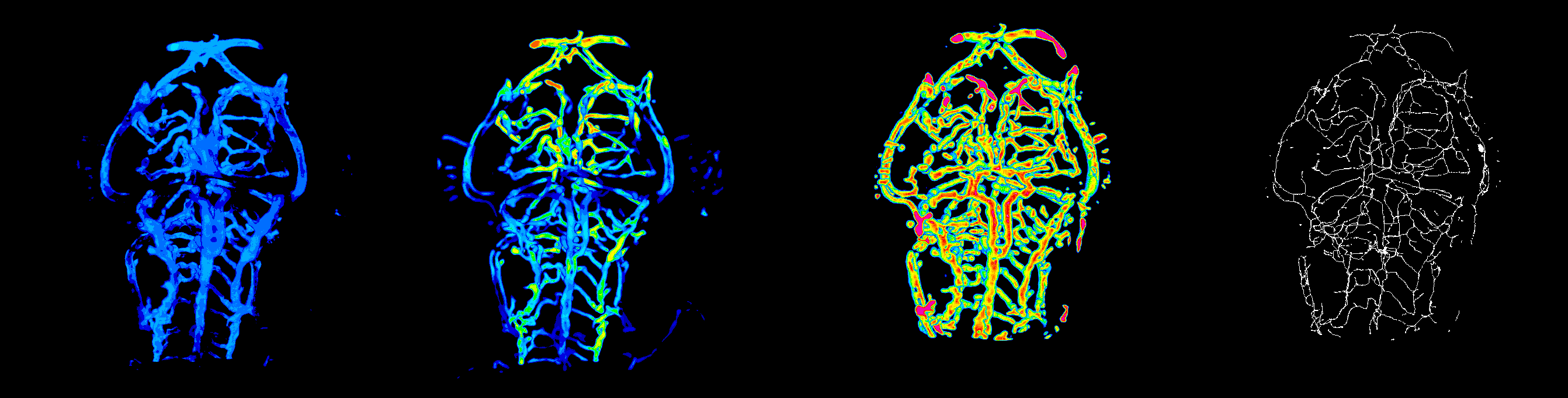
#ZVQ - 3D Quantification of the Zebrafish Brain Vasculature
Vascular diseases are the leading cause of death world-wide. Understanding vascular development and disease is crucial in identifying future options for prevention and treatment.
Detailed insights into vascular development in vivo and 3D can be achieved using zebrafish transgenic lines and light sheet fluorescence microscopy.
However, robust quantification of the zebrafish cerebral vasculature in 3D remains a major challenge due to the lack of tailored analysis tools.
For her PhD at the University of Sheffield, Elisabeth developed image analysis approaches to understand, quantify, and describe the 3D zebrafish brain vascular architecture.
The developed analysis pipeline included
-
image understanding and characterisation: Contrast-to-Noise ratio, motion artefacts/correction, data scales.
-
enhancement and segmentation: pre-processing filters (general filters, Sato, Frangi), segmentation (thresholding, advanced methods such as level set), segmentation validation.
-
registration: landmark-based and automatic.
-
quantification: volume, surface, density, length, branching points, diameter, complexity; global scale and sub-regions.
Together, her work allows for more a comprehensive descriptions of vascular development and the identification of the role of genetic or chemical components.
Publications Derived From This Research
Elisabeth C. Kugler*, James Frost, Vishmi Silva, Karen Plant, Karishma Chhabria, Tim J. A. Chico , Paul A. Armitage*,
Zebrafish vascular quantification: a tool for quantification of three-dimensional zebrafish cerebrovascular architecture by automated image analysis, Development (2022) 149 (3): dev199720. https://doi.org/10.1242/dev.199720
Kugler E.*, Rampun A., Chico T., and Armitage P., Segmentation of the Zebrafish Brain Vasculature from Light Sheet Fluorescence Microscopy Datasets. BioRxiv, 2020
Conference Paper - Kugler E.*, Chico T. and Armitage P., Validating Segmentation of the Zebrafish Vasculature, In: Zheng Y., Williams B., Chen K. (eds) Medical Image Understanding and Analysis. MIUA 2019. Communications in Computer and Information Science, vol 1065. Springer, Cham
Journal Article - Kugler E.*, Plant K., Chico T. and Armitage P., Enhancement and Segmentation Workflow for the Developing Zebrafish Vasculature, Journal of Imaging, MDPI, Vol. 5, Issue 1. 2019
Conference Paper - Kugler E.*, Chico T. and Paul A., Image Analysis in Light Sheet Fluorescence Microscopy Images of Transgenic Zebrafish Vascular Development, In: Nixon, M., Mahmoodi, S. and Zwiggelaar, R., (eds.) Medical Image Understanding and Analysis. MIUA 2018, 09-11 Jul 2018, Southampton, UK. Communications in Computer and Information Science, 894 . Springer Nature Switzerland AG , pp. 343-353. ISBN 9783319959207